问题描述
base-R函数all.vars()返回表达式中的所有名称。例如:
> all.vars( ~ e == M * c^2 )
[1] "e" "M" "c"
有一个不适合使用的R运算符。在许多不使用非标准评估和rlang及其类似函数的人编写的表达式中,名称将是变量的名称。但是,如果这些表达式包含对$的调用,则右侧的名称将不是变量,而是索引或列名。 (我知道变量和列名之间的区别可以通过巧妙地使用环境和数据屏蔽来模糊,但这不是重点。)
all.vars()不能忽略$的右侧。是否有任何类似的功能,或者我必须编写自己的expression-walker?基本上,我想要一个函数,如果传递表达式
a $ b + c $ d
将返回“ a”和“ c”。
请求原因
罗兰,您最好建议我解释一下我为什么要这么做。我经常使用向量化,因为这是我在非常大的数据集上进行计算时获得足够速度的唯一方法。因此,我的代码中的大部分都包含以下内容:
cond <- ¢ A logical vector of 500,000 elements ¢
v1 ¢ (and v2 etc. ) ¢ <- ¢ Numerical or string vectors of the same length ¢
result_size <- length( cond )
result <- rep( NA,result_size )
result[ cond ] <- f( v1[ cond ],v3[ cond ],v4[ cond ],v7[ cond ],v9[ cond ],v10[ cond ]
)
result[ ! cond ] <- g( v2[ ! cond ],v3[ ! cond ],v4[ ! cond ],v5[ ! cond ],v6[ ! cond ],v8[ ! cond ],v10[ ! cond ]
)
我认为这就是R专家所谓的拆分工作流程。按条件拆分数据,分别处理每个组,合并结果。
这种模式正在被抽象为看起来像条件的东西。 (例如,参见RD Tennant的书,编程语言的语义学,以这种意义上的抽象示例以及它们为什么很好。)因此,除了上面容易发生意外的内容外,还应充实可键入和错误的内容。重复索引和部分向量分配,我希望能够编写:
cond <- ¢ A logical vector of 500,000 elements ¢
v1 ¢ (and v2 etc. ) ¢ <- ¢ Numerical or string vectors of the same length ¢
result <- splivif( cond,f( v1,v3,v4,v7,v9,v10 ),g( v2,v5,v6,v8,v10 )
)
splivif()应该被解释为另一种条件函数,就像if(),if_else(),ifelse()和fifelse()一样,并且我可能还不知道的其他六个人。它将以与那些函数相同的方式隐藏一些巧妙的细节:即评估其条件,根据该条件将“ then”和“ else”分支中提到的所有变量拆分为子向量,在这些子向量上调用每个分支,然后结合结果。
这种功能实际上是我经常实现和使用的功能。如上所述,它首先评估其状况。然后,它在“ then”和“ else”表达式中扫描变量。它评估那些被认为是正确长度的向量的向量。然后,它仅选择每个值的第cond个元素,并将它们在新环境中绑定到原始变量的名称。
因此,到本阶段结束时,我们有了一个新的婴儿环境E,其中名称“ v1”绑定到V1[ cond ],其中V1是原始v1。在E中,名称“ v2”到“ v10”的绑定也类似。
splivif()然后在E中计算其“ then”和“ else”表达式,并将结果合并为结果向量。
我对all.vars()的需求是在“ then”和“ else”表达式中扫描变量。我使用嵌套的命名列表来存储控制我的计算的值。因此,这些表达式对这些列表的元素的引用很少,例如Taxogellation $ IgnoreRepeats和Taxogellation $ DoInnerSplines。我问的一个小问题是,all.vars()然后会错误地返回“ IgnoreRepeats”和“ DoInnerSplines”作为变量的名称,而实际上它们是索引。
代码来说明为什么需要向量化
为响应Roland的评论,我于2020年9月8日添加了本节。它建立了一个样本数据表,代表了50,000个家庭的收入,年龄和健康状况。每个家庭由一个或两个成年人组成。然后,它定义一个函数pension(),该函数计算每个家庭应得的退休金。这与现任政府所提供的都不一样,但是说明了典型的养老金计算的特征。例如,结果通常取决于年龄和健康状况,并且可能取决于收入。这些依赖性为任何此类计算设置了最小的复杂度,因此也使时间最短。
然后,代码将对所有50,000个家庭应用pension()的三种方式进行比较和计时。它们是:通过Tidyverse分组;通过data.table分组;和向量化。后者利用这样的事实,可以将诸如+,|,>和pmax()之类的运算符和函数应用于多个元素的向量,然后在相应的元素。
我的计时结果表明,与矢量化相比,Tidyverse甚至data.table的速度都非常慢。对于50,000个家庭,向量化速度要快40倍!
library( tidyverse )
library( data.table )
library( assertthat )
library( microbenchmark )
library( purrr )
#1) Create sample data
#=====================
# The code in this section makes a table
# of no_of_groups families. Each family has
# one or two adults. Adults are randomly assigned
# an income,between 0 and 30,000 pounds;
# an age,between 18 and 99,and a health
# indicator. Each family also gets an integer
# ID. Each adult also gets a number indicating
# whether they are adult 1 or adult 2.
#
# The sections following this will define
# a function for calculating families'
# pensions. My code will apply it in three
# ways,and time each one. These are: by
# grouping using the Tidyverse; by grouping
# using data.table; and by vectorisation.
# This shows that the Tidyverse and data.table
# are both woefully inefficient compared with
# vectorisation. For 5,000 families,the
# Tidyverse takes 2.5 seconds and data.table
# 2 seconds. Vectorisation takes a mere 50
# milliseconds,40 times as fast.
#
no_of_groups <- 5000
group_sizes <- sample( c(1,2),no_of_groups,replace=TRUE )
ids <- 1:no_of_groups
data <- tibble( fam_id=map2( ids,group_sizes,rep ) %>% unlist() )
data <-
data %>%
group_by( fam_id ) %>%
mutate( ad_no = seq_along( fam_id ),two_people = length( ad_no ) == 2
) %>%
ungroup()
data $ income <- runif( nrow( data ),1 ) * 30000
data $ age <- sample( 18:99,nrow( data ),replace=TRUE )
data $ bad_health <- sample( c(T,F),replace=TRUE,prob=c(0.1,0.9) )
#2) Function to calculate pension on single family
#=================================================
# two_people is true if the family has two
# people,otherwise false.
# ad1_inc and ad2_inc are the incomes,in
# pounds per year. ad2_inc is NA if there is
# only one person.
# Similarly,ad1_age and ad2_age are ages.
# And ad1_bad_health and ad2_bad_health are
# Booleans indicating whether the person
# has bad health.
# The result is the pension the Government
# gives the family,in pounds per week.
# This is NOT meant to be the same as in any
# existing country's social-security system,# but exemplifies the kinds of calculation
# such a function needs to do. On our data,# these will be called several hundred
# thousand times.
#
pension <- function( two_people,ad1_inc,ad2_inc,ad1_age,ad2_age,ad1_bad_health,ad2_bad_health
)
{
max_age <-
ifelse( two_people,pmax( ad1_age,ad2_age ),ad1_age
)
income <-
ifelse( two_people,ad1_inc + ad2_inc,ad1_inc
)
bad_health <-
ifelse( two_people,ad1_bad_health | ad2_bad_health,ad1_bad_health
)
pension_level <-
case_when( income > 50000 | max_age < 65 ~ "None",max_age > 80 | bad_health ~ "High",max_age >= 65 ~ "Normal"
)
pension <-
case_when( pension_level == "High" ~ 200.00,pension_level == "Normal" ~ 150.00,pension_level == "None" ~ 0
)
pension
}
#3) Check it works
#=================
pension( F,40000,NA,75,F,NA )
# 150.
pension( T,20000,F )
# 150.
pension( F,60000,NA )
# 0,because of high income.
pension( T,30000,F )
# 0,because of high income.
pension( F,50,because of low age.
pension( T,T )
# 200,because of bad health.
#4) Function to calculate all pensions using Tidyverse group-by
#==============================================================
pension_over_all_TV <- function( data )
{
results <-
data %>%
group_by( fam_id ) %>%
group_map( ~ {
assert_that( nrow( .x ) %in% c( 1,2 ) )
two_people <- .x $ two_people[[ 1 ]]
pension( two_people,.x $ income[[ 1 ]],ifelse( two_people,.x $ income[[ 2 ]],NA ),.x $ age [[ 1 ]],.x $ age[[ 2 ]],.x $ bad_health[[ 1 ]],.x $ bad_health[[ 2 ]],NA )
)
}
)
#
# A vector of pension values,one per family.
results
}
#5) Try it and time it
#=====================
pensions_TV <- pension_over_all_TV( data )
#
# Pensions as calculated by Tidyverse grouping.
res <- microbenchmark( pension_over_all_TV( data ),times=3 )
print( res )
#
# Time it. Mean is 2.5 seconds:
# Unit: seconds
# expr min lq mean median uq max neval
# pension_over_all_TV(data) 2.533073 2.565714 2.584183 2.598356 2.609738 2.621121 3
#6) Function to calculate all pensions using data.table group-by
#===============================================================
pension_over_all_DT <- function( data )
{
# The function that data.table must apply
# to each group.
#
f <- function( group )
{
assert_that( nrow( group ) %in% c( 1,2 ) )
two_people <- group $ two_people[[ 1 ]]
pension( two_people,group $ income[[ 1 ]],group $ income[[ 2 ]],group $ age [[ 1 ]],group $ age[[ 2 ]],group $ bad_health[[ 1 ]],group $ bad_health[[ 2 ]],NA )
)
}
data <- as.data.table( data )
results <-
data[,f( .SD ),by=c( "fam_id" )
]
#
# A table with a V1 column containing one
# pension value per family.
results
}
#7) Try it and time it
#=====================
pensions_DT <- pension_over_all_DT( data )
#
# Pensions as calculated by data.table grouping.
assert_that( are_equal( unlist( pensions_TV ),pensions_DT $ V1 ) )
#
# Making allowance for the slightly different
# formats of the results returned by group_map()
# and data.table's grouped operations,check
# that the numbers are the same.
res <- microbenchmark( pension_over_all_DT( data ),times=3 )
print( res )
#
# Time it. Mean is 2 seconds:
# Unit: seconds
# expr min lq mean median uq max neval
# pension_over_all_DT(data) 1.824391 1.950273 2.155805 2.076154 2.321512 2.56687 3
#8) Function to calculate all pensions using vectorisation
#=========================================================
# This applies pension() to data by using vectorisation.
# It widens data into a table wherein each column is
# a vector corresponding to one of pension()'s arguments.
# It then calls exec() to apply pension() to these
# vectors. I had deliberately written pension() so that
# it would work on vector arguments with more than one
# element.
#
pension_over_all_Vect <- function( data )
{
data_widened <-
pivot_wider( data,names_from = "ad_no",names_prefix = "ad",values_from = all_of( c("income","age","bad_health") )
) %>%
rename( ad1_inc="income_ad1",ad2_inc="income_ad2",ad1_age="age_ad1",ad2_age="age_ad2",ad1_bad_health="bad_health_ad1",ad2_bad_health="bad_health_ad2"
) %>%
select( -fam_id )
#
# A table with one row for each family,and one
# column for each of pension()'s arguments.
results <- exec( pension,!!! as.list( data_widened ) )
#
# A vector of results: one pension value for
# each family.
results
}
#9) Try it and time it
#=====================
pensions_Vect <- pension_over_all_Vect( data )
#
# Returns a list of plausible-looking results.
assert_that( are_equal( unlist( pensions_TV ),pensions_Vect ) )
assert_that( are_equal( unlist( pensions_DT $ V1 ),pensions_Vect ) )
#
# Check that this is equal to the previously-
# calculated results.
res <- microbenchmark( pension_over_all_Vect( data ),times=3 )
print( res )
#
# Time it. The mean is 50 milliseconds.
# Unit: milliseconds
# expr min lq mean median uq max neval
# pension_over_all_Vect(data) 35.7834 45.23245 50.8431 54.6815 58.37295 62.0644 3
解决方法
您应该解释为什么需要这样做以及其他形式的非标准评估会发生什么。对于您的实际问题,可能会有更好的解决方案。
我会迅速将$替换为[[:
replace_dollar <- function(expr) {
if (!is.language(expr) || length(expr) == 1L) return(expr)
if (expr[[1]] == quote(`$`)) {
expr[[1]] <- quote(`[[`)
expr[[3]] <- as.character(expr[[3]])
} else {
for (i in seq_along(expr)[-1])
expr[[i]] <- replace_dollar(expr[[i]])
}
expr
}
expr <- quote(a $ b + c $ d)
replace_dollar(expr)
# a[["b"]] + c[["d"]]
all.vars(replace_dollar(expr))
#[1] "a" "c"
请注意,根据文档:
x$name等效于x[["name",exact = FALSE]]。
我认为您在这里不关心部分名称匹配,因为您只想传递给all.vars。
您可以使用all.vars选项输出包括$运算符在内的整个结构,并从列表中删除$运算符的第二个参数:
test <- ~a$b+c$d
all <- all.vars(test,functions = T,unique = F)
all
#> [1] "~" "+" "$" "a" "b" "$" "c" "d"
to_remove <- all[c(F,F,all == "$")]
to_remove
#> [1] "b" "d"
vars <- all.vars(test)
vars
#> [1] "a" "b" "c" "d"
vars[!vars %in% to_remove]
#> [1] "a" "c"
<sup>Created on 2020-08-25 by the [reprex package](https://reprex.tidyverse.org) (v0.3.0)</sup>
或作为功能:
all.vars.new <- function(e) {
all <- all.vars(e,unique = F)
cols <- all[c(F,all == "$")]
vars <- all.vars(e)
vars[!vars %in% cols]
}

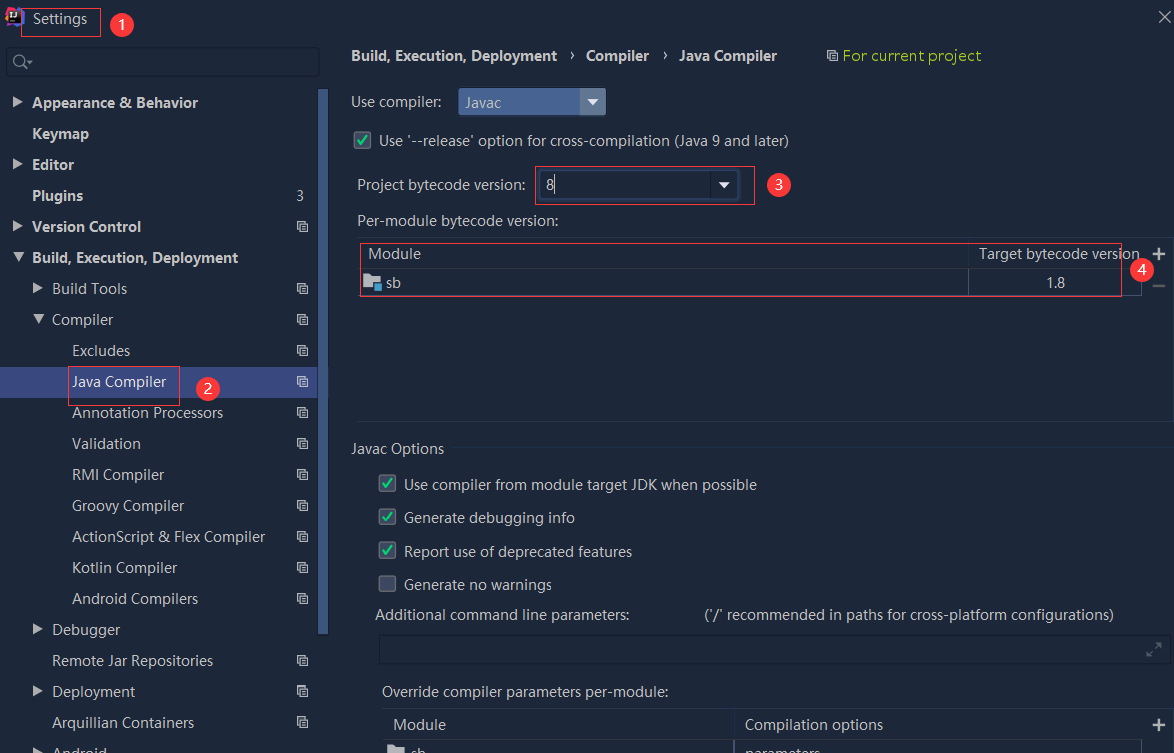 依赖报错 idea导入项目后依赖报错,解决方案:https://blog....
依赖报错 idea导入项目后依赖报错,解决方案:https://blog....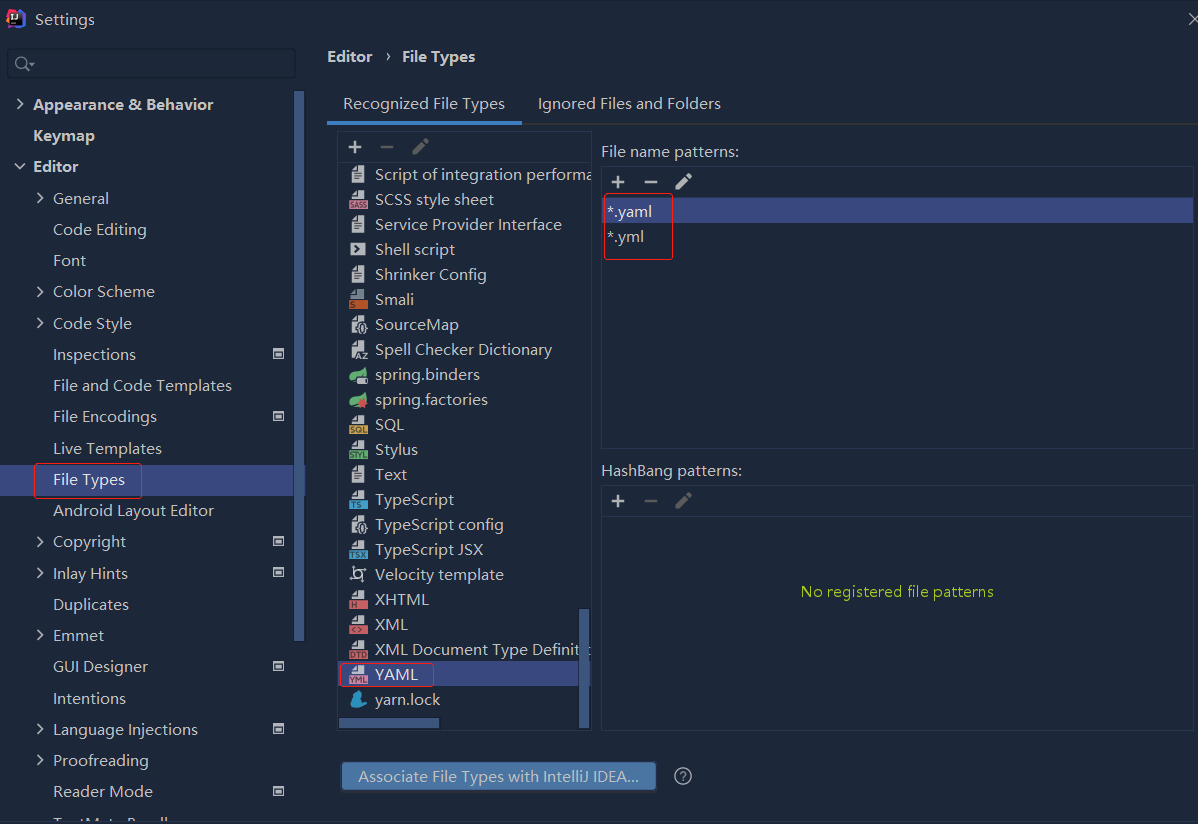
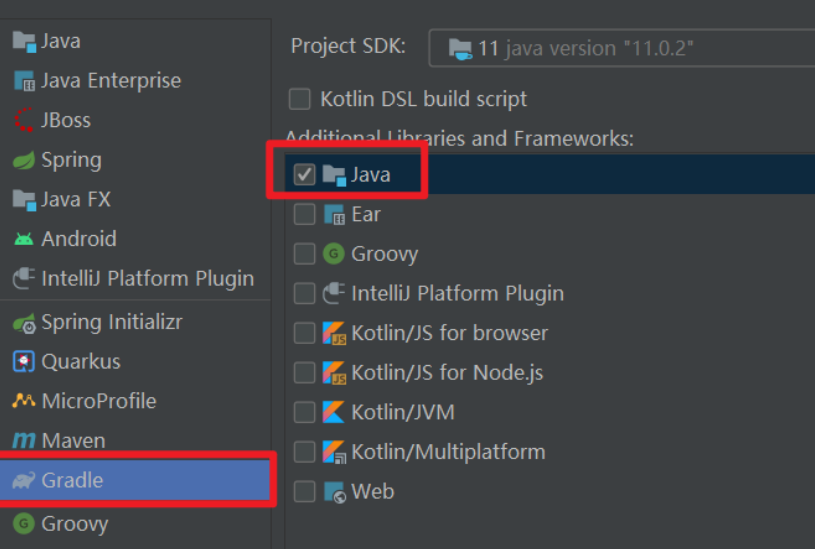 错误1:gradle项目控制台输出为乱码 # 解决方案:https://bl...
错误1:gradle项目控制台输出为乱码 # 解决方案:https://bl...